Protein-coding gene in the species Homo sapiens
PTCH1 Identifiers Aliases PTCH1 , BCNS, HPE7, NBCCS, PTC, PTC1, PTCH, PTCH11, patched 1External IDs OMIM: 601309; MGI: 105373; HomoloGene: 223; GeneCards: PTCH1; OMA:PTCH1 - orthologs Gene location (Human) Chr. Chromosome 9 (human)[1] Band 9q22.32 Start 95,442,980 bp[1] End 95,517,057 bp[1]
Gene location (Mouse) Chr. Chromosome 13 (mouse)[2] Band 13 B3|13 32.8 cM Start 63,656,142 bp[2] End 63,721,412 bp[2]
RNA expression pattern Bgee Human Mouse (ortholog) Top expressed in tibia spinal ganglia trigeminal ganglion sural nerve pylorus sperm retinal pigment epithelium cardia internal globus pallidus endometrium
Top expressed in upper jaw Gonadal ridge tooth molar vas deferens migratory enteric neural crest cell lobe of cerebellum sciatic nerve hair follicle dermis
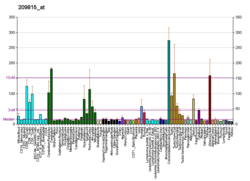
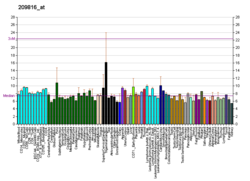
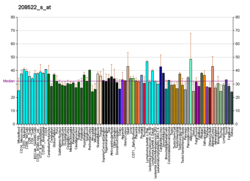
More reference expression data
BioGPS More reference expression data
Gene ontology Molecular function
heparin binding
smoothened binding
cholesterol binding
patched binding
protein binding
hedgehog receptor activity
hedgehog family protein binding
cyclin binding
protein-containing complex binding Cellular component
integral component of membrane
endocytic vesicle membrane
dendritic growth cone
Golgi apparatus
intracellular membrane-bounded organelle
postsynaptic density
membrane
plasma membrane
axonal growth cone
midbody
ciliary membrane
perinuclear region of cytoplasm
caveola
cilium
nucleus Biological process
pattern specification process
renal system development
neural plate axis specification
response to estradiol
limb morphogenesis
negative regulation of smoothened signaling pathway
response to organic cyclic compound
response to retinoic acid
glucose homeostasis
embryonic organ development
cell proliferation involved in metanephros development
negative regulation of multicellular organism growth
protein processing
response to mechanical stimulus
heart morphogenesis
epidermal cell fate specification
in utero embryonic development
negative regulation of transcription by RNA polymerase II
hindlimb morphogenesis
negative regulation of DNA-binding transcription factor activity
negative regulation of osteoblast differentiation
mammary gland epithelial cell differentiation
somite development
cellular response to cholesterol
regulation of protein localization
positive regulation of transcription, DNA-templated
mammary gland duct morphogenesis
mammary gland development
neural tube patterning
regulation of smoothened signaling pathway
branching involved in ureteric bud morphogenesis
neural tube closure
brain development
keratinocyte proliferation
negative regulation of cell division
smoothened signaling pathway involved in dorsal/ventral neural tube patterning
regulation of cell population proliferation
embryonic limb morphogenesis
positive regulation of cholesterol efflux
regulation of mitotic cell cycle
negative regulation of epithelial cell proliferation
epidermis development
animal organ morphogenesis
cell differentiation involved in kidney development
cell fate determination
response to chlorate
regulation of growth
smoothened signaling pathway
dorsal/ventral neural tube patterning
spinal cord motor neuron differentiation
negative regulation of transcription, DNA-templated
positive regulation of epidermal cell differentiation
dorsal/ventral pattern formation
neural tube formation
signal transduction
negative regulation of cell population proliferation
pharyngeal system development
prostate gland development
commissural neuron axon guidance
protein localization to plasma membrane
liver regeneration Sources:Amigo / QuickGO
Orthologs Species Human Mouse Entrez Ensembl UniProt RefSeq (mRNA) NM_000264 NM_001083606
RefSeq (protein) NP_000255 NP_001077075
Location (UCSC) Chr 9: 95.44 – 95.52 Mb Chr 13: 63.66 – 63.72 Mb PubMed search [3] [4]
Wikidata View/Edit Human View/Edit Mouse
Protein patched homolog 1 is a protein that is the member of the patched family and in humans is encoded by the PTCH1 gene.[5] [6]
^ a b c GRCh38: Ensembl release 89: ENSG00000185920 – Ensembl, May 2017
^ a b c GRCm38: Ensembl release 89: ENSMUSG00000021466 – Ensembl, May 2017
^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine . ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine . ^ Johnson RL, Rothman AL, Xie J, Goodrich LV, Bare JW, Bonifas JM, Quinn AG, Myers RM, Cox DR, Epstein EH, Scott MP (Aug 1996). "Human homolog of patched, a candidate gene for the basal cell nevus syndrome". Science . 272 (5268): 1668–71. Bibcode:1996Sci...272.1668J. doi:10.1126/science.272.5268.1668. PMID 8658145. S2CID 9160210. ^ "Entrez Gene: PTCH1 patched homolog 1 (Drosophila)".
Last Update: 2023-12-23T10:59:30Z
the member of the patched family and in humans is encoded by the PTCH1 gene. PTCH1 is a member of the patched gene family and is the receptor for sonic...
Word Count : 1629
Last Update: 2024-04-25T22:30:14Z
it binds to the Patched-1 (PTCH1 ) receptor (Process "3" on Figure 5, the blue molecule). In the absence of ligand, PTCH1 inhibits Smoothened (SMO), a...
Word Count : 6895
Last Update: 2024-04-26T22:51:04Z
syndrome as well as Turcot syndrome. Recurrent mutations in the genes CTNNB1, PTCH1 , MLL2, SMARCA4, DDX3X, CTDNEP1, KDM6A, and TBR1 were identified in individuals...
Word Count : 3455
Last Update: 2023-12-04T01:11:37Z
Patched1 (PTCH1 ). In the absence of Shh, PTCH1 inhibits the activity of Smoothened (SMO), another transmembrane protein. Upon the inactivation of PTCH1 by Shh...
Word Count : 3864
Last Update: 2024-06-05T19:18:48Z
A molecular factor involved in the disease process is mutation in gene PTCH1 that plays an important role in the Sonic hedgehog signaling pathway. Diagnosis...
Word Count : 6427
Last Update: 2023-12-03T19:46:27Z
translocated into the nucleus where they activate target genes such at PTCH1 and Engrailed. Drosophila melanogaster has an unusual developmental system...
Word Count : 1848
Last Update: 2024-03-18T09:44:23Z
NDUFB11 NHS OTX2 PAX2 PAX6 PDE6D PIGL POLR1C POLR1D PORCN PQBP1 PRSS56 PTCH1 RAB3GAP1 RAB3GAP2 RARB RAX RBP4 RPGRIP1L SALL1 SALL2 SALL4 SCLT1 SEMA3E...
Word Count : 2173
Last Update: 2022-03-28T03:03:27Z
and rhabdomyosarcoma. Hereditary mutations in the human patched homolog PTCH1 cause autosomal dominant Gorlin syndrome, which consists of overgrowth and...
Word Count : 442
Last Update: 2024-04-28T09:12:54Z
location missing publisher (link) "PATCHED, DROSOPHILA, HOMOLOG OF, 1; PTCH1 ". OMIM. Ren C, Amm HM, DeVilliers P, Wu Y, Deatherage JR, Liu Z, MacDougall...
Word Count : 2235
Last Update: 2024-06-17T03:39:46Z
oncogenes), a co-receptor for Shh that influences Shh signaling through Ptch1 , seems to mediate this repulsion, as it is only on growth cones coming from...
Word Count : 3521
Last Update: 2022-06-15T15:45:04Z
Shh regulates Gli processing through two proteins, Ptch1 and Smo. When Shh is not active, Ptch1 is responsible for suppressing the pathway through the...
Word Count : 1961
Last Update: 2024-06-22T15:36:01Z
Barakat syndrome (HDR syndrome) 146255 GATA3 Basal cell nevus syndrome 109400 PTCH1 , PTCH2, SUFU Branchio‐oculo‐facial syndrome 113620 TFAP2A C syndrome (Opitz...
Word Count : 2499
Last Update: 2023-08-13T12:03:37Z
gene localizes in the cytoplasm and activates patched Drosophila homolog (PTCH1 ) gene expression. It is also thought to play a role during embryogenesis...
Word Count : 1196
Last Update: 2024-04-11T01:50:05Z
mitochondrial processing alpha subunit PRUNE2: protein prune homolog 2 PTCH1 : protein patched 1 transmembrane receptor protein RABGAP1: RAB GTPase activating...
Word Count : 1594
Last Update: 2024-01-05T23:00:30Z
of basal cell carcinoma. Basal cell carcinoma is induced by mutations in PTCH1 , a tumour-suppressor protein, leading to uncontrollable cell growth. In...
Word Count : 2371
Last Update: 2024-03-18T09:46:51Z
malformation syndrome (HBMS) 14q32 AD Hemifacial microsomia SIX3, SHH, PTCH1 , GLI2 AD Holoprosencephaly types 1, 2, 3, 7, 9 IKBKG XLD Incontinentia pigmenti...
Word Count : 538
Last Update: 2023-10-17T20:24:43Z
Albert Pinhasov, et al., "Differences in RNA and microRNA Expression Between PTCH1 - and SUFU-mutated Medulloblastoma", Cancer Genomics & Proteomics Vol. 18...
Word Count : 1120
Last Update: 2022-05-06T16:23:42Z
development and cell formation. Matthew P. Scott has identified the human homolog PTCH1 as a key tumor suppressor gene for the Hedgehog signaling pathway and the...
Word Count : 1404
Last Update: 2023-12-03T02:05:31Z
Roychoudhury S, Panda CK (2008). "Alterations in candidate genes PHF2, FANCC, PTCH1 and XPA at chromosomal 9q22.3 region: pathological significance in early-...
Word Count : 205
Last Update: 2024-05-01T06:40:33Z
IRF6, MAFB (gene), MMP3, MSX1, MSX2 (Msh homeobox 2), MSX3, PAX7, PDGFC, PTCH1 , SATB2, SOX9, SUMO1 (Small ubiquitin-related modifier 1), TBX22, TCOF (Treacle...
Word Count : 610
Last Update: 2023-12-02T22:03:05Z
Carcinoma) UNC5 receptors (UNC5A, UNC5B, UNC5C, UNC5D) Neogenin p75NTR Ptch1 CDON PLXND1 RET TrkA TrkC EphA4 c-Met Insulin receptor IR Insulin-like growth...
Word Count : 1432
Last Update: 2024-05-15T05:51:09Z
differentiation. Stellate cells express smoothened (Smo) and patched-1 (Ptch1 ) proteins, which are significant features of the hedgehog receptor system...
Word Count : 2479
Last Update: 2024-05-09T15:43:25Z
605462; PTCH1 Basal cell carcinoma, somatic; 605462; PTCH2 Basal cell carcinoma, somatic; 605462; RASA1 Basal cell nevus syndrome; 109400; PTCH1 Basal ganglia...
Word Count : 18877
 Global Information
Global Information