 Global Information
Global InformationPii nitrogen regulatory proteins information
| Nitrogen regulatory protein P-II | |||||||||
|---|---|---|---|---|---|---|---|---|---|
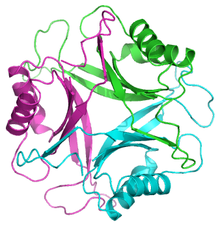 GlnB protein from E.coli. PDB 2pii[1] The GlnB protein in E. coli is 68% identical to GlnK. | |||||||||
| Identifiers | |||||||||
| Symbol | P-II | ||||||||
| Pfam | PF00543 | ||||||||
| Pfam clan | CL0089 | ||||||||
| InterPro | IPR002187 | ||||||||
| SMART | SM00938 | ||||||||
| PROSITE | PDOC00439 | ||||||||
| SCOP2 | 1pil / SCOPe / SUPFAM | ||||||||
| |||||||||
The PII family comprises a group of widely distributed signal transduction proteins found in nearly all Bacteria and also present in Archaea and in the chloroplasts of Algae and plants.[2] PII form barrel-like homotrimers with a flexible loop, namely T-loop, emerging from each subunit.[2] PII proteins have extraordinary sensory properties; they can exist in a vast range of structural status accordingly to the levels of ATP, ADP and 2-oxogluratate. These metabolites interact allosterically with PII in three conserved binding sites located in the lateral cavity between each PII subunit. ATP and ADP bind competitively to the nucleotide binding whereas the 2-oxoglutarate only interacts with PII in the presence of MgATP.[3]
In Proteobacteria, PII proteins are also subject to a cycle of reversible posttranslational modification (Huergo et al., 2013). Under a low nitrogen regime, the low intracellular glutamine level triggers the uridylyl-transferase activity of the bi-functional GlnD enzyme promoting the uridylylation of a conserved Tyr-51 located at the top the PII T-loop. Conversely, under a high nitrogen regime, accumulation of intracellular glutamine triggers the uridylyl-removing activity of GlnD and PII accumulates in its non-modified form (Huergo et al., 2013).
The ability of PII to sense important metabolites and integrate signals deriving from energy status (ATP and ADP ratio), carbon (2-oxogluratate) and nitrogen (glutamine and 2-oxoglutarate) levels were capitalized during evolution such that PII can act as a dissociable regulatory subunit of a range of transporters, transcriptional regulators and enzymes.[4]
- ^ Carr PD, Cheah E, Suffolk PM, Vasudevan SG, Dixon NE, Ollis DL (January 1996). "X-ray structure of the signal transduction protein from Escherichia coli at 1.9 A". Acta Crystallographica Section D. 52 (Pt 1): 93–104. Bibcode:1996AcCrD..52...93C. doi:10.1107/S0907444995007293. PMID 15299730.
- ^ a b Huergo LF, Chandra G, Merrick M (March 2013). "P(II) signal transduction proteins: nitrogen regulation and beyond". FEMS Microbiology Reviews. 37 (2): 251–83. doi:10.1111/j.1574-6976.2012.00351.x. PMID 22861350.
- ^ Huergo LF, Pedrosa FO, Muller-Santos M, Chubatsu LS, Monteiro RA, Merrick M, Souza EM (January 2012). "PII signal transduction proteins: pivotal players in post-translational control of nitrogenase activity". Microbiology. 158 (Pt 1): 176–90. doi:10.1099/mic.0.049783-0. PMID 22210804.
- ^ Huergo LF, Dixon R (December 2015). "The Emergence of 2-Oxoglutarate as a Master Regulator Metabolite". Microbiology and Molecular Biology Reviews. 79 (4): 419–35. doi:10.1128/MMBR.00038-15. PMC 4651028. PMID 26424716.