 Global Information
Global InformationPYLIS downstream sequence information
| Pyrrolysine insertion sequence 1 | |
|---|---|
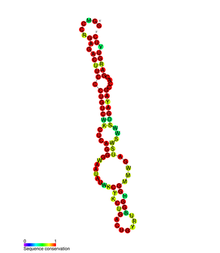 Predicted secondary structure of PYLIS 1 RNA. | |
| Identifiers | |
| Symbol | PYLIS 1 |
| Rfam | RF01982 |
| Other data | |
| RNA type | Cis-regulatory element |
| Domain(s) | Archaea |
| PDB structures | PDBe |
In biology, the PYLIS downstream sequence (PYLIS: pyrrolysine insertion sequence) is a stem-loop structure that appears on some mRNA sequences. This structural motif was previously thought to cause the UAG (amber) stop codon to be translated to the amino acid pyrrolysine instead of ending the protein translation.[1][2][3] However, it has been shown that PYLIS has no effect upon the efficiency of the UAG suppression,[4] hence even its name is, in fact, incorrect.
- ^ Namy O, Rousset JP, Napthine S, Brierley I (2004). "Reprogrammed genetic decoding in cellular gene expression". Molecular Cell. 13 (2): 157–168. doi:10.1016/S1097-2765(04)00031-0. PMID 14759362.
- ^ Théobald-Dietrich A, Giegé R, Rudinger-Thirion J (2005). "Evidence for the existence in mRNAs of a hairpin element responsible for ribosome dependent pyrrolysine insertion into proteins". Biochimie. 87 (9–10): 813–817. doi:10.1016/j.biochi.2005.03.006. PMID 16164991.
- ^ Zhang Y, Baranov PV, Atkins JF, Gladyshev VN (May 2005). "Pyrrolysine and selenocysteine use dissimilar decoding strategies". J. Biol. Chem. 280 (21): 20740–20751. doi:10.1074/jbc.M501458200. PMID 15788401.
- ^ Namy, Olivier; Zhou, Yu; Gundllapalli, Sarath; Polycarpo, Carla R.; Denise, Alain; Rousset, Jean-Pierre; Söll, Dieter; Ambrogelly, Alexandre (November 2007). "Adding pyrrolysine to the Escherichia coli genetic code". FEBS Letters. 581 (27): 5282–5288. doi:10.1016/j.febslet.2007.10.022. PMID 17967457.